## First check for the required packages, install if needed, and load the libraries.
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("sangerseqR")
remotes::install_github("ropensci/bold")
remotes::install_github("ropensci/taxize")
if (!require("pacman")) install.packages("pacman")
pacman::p_load(maps, ggplot2, countrycode, rgbif, data.table, raster, mapproj, sf)Coverage by climatic zones
This notebook will generate plots of geographic coverage of barcodes available on BOLD, in relation to climatic zones, and for several genetic markers. It also assesses contributions to BOLD by Project ID.
We trim dataframe outputs to the first two rows for demonstration, but full dataframes will be generated and stored in the R environment when running the code from source.
Figure 1. Read in combtab dataset of global barcode data from BOLD and summarize.
combtab <- read.csv("../data/BoldPhyla_to_Families_combtab_v4.csv")
#View(combtab_20240624)
combtab_2 <- (unique(combtab$family_name))
head(combtab_2, 2)[1] "Acanthaceae" "Achariaceae"head(unique(combtab$family_name), 2)[1] "Acanthaceae" "Achariaceae"combtab_subset_barcodes <- combtab %>% subset(multi != "---" & multi != "")
head(combtab_subset_barcodes, 2) X.1 X processid institution_storing phylum_taxID
1 1 1 API120-12 Sri Ramaswamy Memorial University 12
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
1 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
1 Acanthaceae NA 415894 Peristrophe
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
species_taxID species_name subspecies_taxID subspecies_name
1 494465 Peristrophe bicalyculata NA
2 632816 Ruellia inflata NA
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
1 NA NA NA NA NA
2 NA NA NA NA NA
elev country province_state region sector exactsite rbcL matK trnL ITS2
1 NA rbcL
2 NA Brazil Para rbcL matK ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS
1 rbcL---
2 rbcL-matK--ITS combtab_split_barcodes <- strsplit(combtab$multi, "-")
head(combtab_split_barcodes, 2)[[1]]
[1] "rbcL" "" ""
[[2]]
[1] "rbcL" "matK" "" "ITS" head(unlist(combtab_split_barcodes)[which(unlist(combtab_split_barcodes)!="")], 2)[1] "rbcL" "rbcL"length(which(unlist(combtab_split_barcodes)!=""))[1] 372460# Convert blank cells to NA
combtab[combtab == ""] <- NA
# Count the number of barcodes per specimen
combtab$barcode_count <- rowSums(!is.na(combtab[, c("rbcL", "matK", "trnL", "ITS2")]))
# Calculate the total number of barcodes across all rows
total_barcodes <- sum(combtab$barcode_count)
# Display the results
#View(combtab)
total_barcodes[1] 373582# write combtab
combtab <- combtab
write.csv(combtab, "../data/combtab.csv")
# write out combtab here for supplementary/supporting information
length(unique(combtab$species_name))[1] 102887length(unique(combtab$country))[1] 208length(unique(combtab$family_name))[1] 651length(unique(combtab$family_name))[1] 651# calculate % north of tropic of cancer, south of tropic of capricorn, and between the two
combtab_north_of_tropic_of_cancer <- combtab %>% subset(lat > 23.4)
head(combtab_north_of_tropic_of_cancer, 2) X.1 X processid institution_storing phylum_taxID phylum_name
106 152 152 GBVU4228-13 Mined from GenBank, NCBI 12 Magnoliophyta
107 153 153 GBVX3812-15 Mined from GenBank, NCBI 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
106 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
107 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
106 NA <NA> 554651 Clarkeasia 554652
107 NA <NA> 210661 Avicennia 641297
species_name subspecies_taxID subspecies_name collectiondate_start
106 Clarkeasia parviflora NA <NA> NA
107 Avicennia officinalis NA <NA> NA
collectiondate_end lat lon coord_source
106 NA 28.3009 84.1339 Coordinates from country centroid
107 NA 23.5863 81.1730 Coordinates from country centroid
coord_accuracy elev country province_state region sector exactsite rbcL
106 NA NA Nepal <NA> Narayani <NA> <NA> rbcL
107 NA NA India <NA> <NA> <NA> <NA> <NA>
matK trnL ITS2 multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count
106 <NA> <NA> <NA> rbcL--- <NA> <NA> <NA> <NA> 1
107 matK <NA> <NA> -matK-- <NA> <NA> <NA> <NA> 1combtab_south_of_tropic_of_capricorn <- combtab %>% subset(lat < -23.4)
head(combtab_south_of_tropic_of_capricorn, 2) X.1 X processid
48 55 55 GBVU4382-13
52 60 60 KNPA505-09
institution_storing
48 Mined from GenBank, NCBI
52 University of Johannesburg, Department of Botany and Plant Biotechnology
phylum_taxID phylum_name class_taxID class_name order_taxID order_name
48 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
52 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
family_taxID family_name subfamily_taxID subfamily_name genus_taxID
48 148533 Acanthaceae NA <NA> 554845
52 148533 Acanthaceae NA <NA> 205058
genus_name species_taxID species_name subspecies_taxID
48 Petalidium 554846 Petalidium oblongifolium NA
52 Ruspolia 205059 Ruspolia hypocrateriformis NA
subspecies_name collectiondate_start collectiondate_end lat lon
48 <NA> NA NA -28.5536 24.7525
52 <NA> NA NA -23.9440 31.7250
coord_source coord_accuracy elev country
48 Coordinates from country centroid NA NA South Africa
52 <NA> NA 208 South Africa
province_state region sector
48 <NA> <NA> <NA>
52 Limpopo <NA> Kruger National Park
exactsite rbcL matK trnL ITS2
48 <NA> rbcL <NA> <NA> <NA>
52 Kruger National Park, Nxanatseni region North, L rbcL matK <NA> <NA>
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count
48 rbcL--- FJ868916 <NA> <NA> <NA> 1
52 rbcL-matK-- KY632577 KY632577 <NA> <NA> 2combtab_between_cancer_capricorn <- combtab %>% subset(lat >= -23.4 & lat <= 23.4)
head(combtab_between_cancer_capricorn, 2) X.1 X processid institution_storing phylum_taxID phylum_name
49 56 56 GENG1220-15 Gujarat Biodiversity Gene Bank 12 Magnoliophyta
50 57 57 GENG1760-16 Gujarat Biodiversity Gene Bank 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
49 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
50 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
49 480072 Thunbergioideae 273749 Thunbergia 317700
50 264310 Acanthoideae 148534 Ruellia NA
species_name subspecies_taxID subspecies_name collectiondate_start
49 Thunbergia grandiflora NA <NA> NA
50 <NA> NA <NA> NA
collectiondate_end lat lon coord_source coord_accuracy elev country
49 NA 22.306 73.278 Garmin etrax 10 NA 130 India
50 NA 21.615 70.505 <NA> NA 532 India
province_state region sector exactsite rbcL matK trnL ITS2 multi
49 Gujarat Vodadara <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
50 Gujarat Junagadh <NA> Etava rbcL <NA> <NA> <NA> rbcL---
gb_rbcL gb_matK gb_trnL gb_ITS barcode_count
49 KX641599 <NA> <NA> <NA> 1
50 KJ535203 <NA> <NA> <NA> 1Assess Project contibutions.
This block will assess the localized sampling projects’ contributions to the overall BOLD database.
# make container column
# overall
head(combtab[grep("UHURU", combtab$processid),], 2) X.1 X processid
59 78 78 UHURU149-14
60 82 82 UHURU394-14
institution_storing
59 Smithsonian Institution, National Museum of Natural History, United States National Herbarium
60 Smithsonian Institution, National Museum of Natural History, United States National Herbarium
phylum_taxID phylum_name class_taxID class_name order_taxID order_name
59 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
60 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
family_taxID family_name subfamily_taxID subfamily_name genus_taxID
59 148533 Acanthaceae NA <NA> 148750
60 148533 Acanthaceae NA <NA> 306313
genus_name species_taxID species_name subspecies_taxID
59 Justicia 591068 Justicia diclipteroides NA
60 Crossandra 659703 Crossandra massaica NA
subspecies_name collectiondate_start collectiondate_end lat lon
59 <NA> NA NA 0.4810 36.8740
60 <NA> NA NA 0.2956 36.8824
coord_source coord_accuracy elev country province_state region sector
59 <NA> NA 1650 Kenya Rift Valley Laikipia <NA>
60 <NA> NA 1650 Kenya Rift Valley Laikipia <NA>
exactsite rbcL matK trnL ITS2 multi gb_rbcL gb_matK
59 Mpala Research Centre rbcL matK trnL <NA> rbcL-matK-trnL- KM255065 KM255065
60 Mpala Research Centre rbcL matK trnL <NA> rbcL-matK-trnL- JX517979 JX517979
gb_trnL gb_ITS barcode_count
59 KM255065 <NA> 3
60 JX517979 <NA> 3combtab$container<-gsub("[0-9.-]", "", combtab$processid)
length(unique(combtab$container))[1] 379sort(table(combtab$container), decreasing=T) #PNG, YNPBP, UHURU
GBITS PGBMA GBVX GBVP JWP GBVA GBVC GBVU GBVE GBVS GBVF GBVW GBVI
70028 10890 8000 6694 6103 5270 4999 4989 4983 4983 4979 4959 4953
GBVM GBVL GBVG GBVH GBVK GBVJ GBVB GBVD GBVT GBVR SAFH GBVO GBVN
4939 4935 4920 4915 4898 4886 4870 4864 4854 4773 4271 3810 3668
GBVY BABRU POWNA EBLF GBVQ CPBOL CRCZ FCA FBPL MHPAD DPC SDH UHURU
3629 3232 3174 3127 2863 2527 2450 2179 2143 2126 1780 1733 1720
GENG EDTOL PNOR KNPA VPSBC FPUK PLOMA BRYCA IDRCG IPSUB TCAFR ABGCO MHPAF
1708 1581 1474 1339 1279 1240 1149 1125 1061 839 780 665 630
MKPCH PLID CANGI GRASS PNG SALIX YNPBP BATT CALAK FSEUS GRDMO AUSG MHPAC
622 590 564 553 551 540 527 520 508 502 498 479 448
TRM MHPAA TDEF SCBI CANGA GCUDG PRTRE KSR ALOAF API FAUCT HOSAM TSA
434 432 429 425 422 384 379 376 350 349 342 341 338
NNOR FPOON KNPB SBIO WABLK TRIOP ALM ANGMD CSPPA VEMSH TMBRY NOVOA GRDTX
337 331 330 324 319 315 314 313 301 297 296 281 280
FGB PHAK GCUBG ITVPL APOSA PSYAF MXBZI TADCR CAATB PIM POWNB CYPAF DBFZP
272 260 256 254 243 233 226 221 214 213 211 205 202
FYNBL VPL RBTEP FOND EGPAM MAPLT PNCA HOLIN POA BPTPS ECU ITVI MARQF
199 197 196 195 188 188 187 184 184 181 179 176 175
TRIKU BRYBC CYAF IASVF HAIDA BBYUK MADA HB MXBTX BRALT SERC CPL CPPB
175 174 173 172 170 164 164 160 160 159 159 157 148
PEPER SAA PLGE BPNP GPAGA KPLA CROT ENDEM COBIN UGEAB WREP MKTRT FLSH
147 143 142 140 138 137 135 135 127 122 120 119 117
MADPL BSCAT RBG PLKEN CAREX MKCRA CONOP NECLE DBMPP SFFIP YHT LWT WAG
111 108 100 99 94 93 92 92 91 89 89 87 85
SASCH CREUA LEDCL LTRBN ZERGT SLNM OSCH RUPL ORPOL LOPHZ POEUA RBPLT VPSEA
84 83 83 83 83 82 81 81 79 78 78 78 76
PLCHA CRBS PCABL BRPLT KIFNE PANFT PLTHU BTAM GRII MEXCO PLNOR VLUZH KIFRH
75 74 74 73 72 70 69 68 68 68 63 63 62
BFAA COBIM DBZN MLTVP MPIN PLCOC RINGV AGOP PLALG SAP AGL ATRIP FSSM
61 60 60 60 60 60 58 55 55 55 54 54 54
PIPER CERF BAMP MIPDG ZPLPP CSU DVHTF DIDA INB SIDA SWFRG BRYUM DBHI
53 51 50 50 49 48 47 46 46 46 46 44 44
CGII MSTH JSUIS HERB IMDB KBGPP NTDB KUDZU MKSAL CACRB IHGTI ITVRT TCHDC
43 43 42 41 40 40 40 39 39 38 36 36 35
KIFMO THY TNAU BBIHP CINCH INDET PDBA DBTS FERNO FMED SYKO PLNIP FAGRB
34 34 34 33 32 31 31 30 29 29 29 28 27
PLNIA LCBAR CACMK FFFCC MELIA MBB PLWEL CRCBS PGLLP ACBRC AGMCV KIFHE ATG
27 26 25 24 23 22 22 21 21 20 20 19 18
BMS FQAGM SMPRS CRCB GTICO TFOAX CRCBT PCUBC RBGEV AGVMK BBOBC USMF SDP
18 18 18 17 17 15 14 14 14 13 13 13 12
CACTB ITICS KIFAT MKOXY STRBS THYME ITVGA MHPAB NELKE PLCHU PLNB PLPAR PTOC
11 11 11 11 11 11 10 10 10 10 10 10 10
UADY AGVRB FAGMK HRUC IMMEX PMRB SABIO BBOTP LIVCA MCT PCCMN PLTOR SEQLD
10 9 9 9 9 9 9 8 8 8 8 8 8
SPBL AUCAS CABAR RHEUM VASCB ALSEU BSRUG DBTB PHSN PLLAM RBJ ABCBF ACCCO
8 7 7 7 7 6 6 6 6 6 6 5 5
BEER BHEPS CBIHP DRAC IIPSA MAHA MPNE MTSS NPOAF PLON QUNAM RSUJO SSAA
5 5 5 5 5 5 5 5 5 5 5 5 5
CBL CSPPB EMPWG RICE BUL COMPL GRJ ITICZ MSHQU MTAT PCYMX RJAS AMBC
4 4 4 4 3 3 3 3 3 3 3 3 2
BCBSL DCCC GYM HIMS HRBC KFRI LGRAS ORIGV PLPET PLSUD SEEWI STRM SUSHI
2 2 2 2 2 2 2 2 2 2 2 2 2
AKD APP ASJ DBALT DLFW DMCSI DRCLS EOCCF GUTB ISO LAARS LADEE LAHAR
1 1 1 1 1 1 1 1 1 1 1 1 1
LAMSK MASMD MCAND MCBRT MCCAL MCGFB MCIII MCJNM MCMAN MCUII MCUKI MCVAD MCYMB
1 1 1 1 1 1 1 1 1 1 1 1 1
PCGM PLBRU PLMUS PLREN SDCCF SIDBG SLAT SLX SPAP SPAR SRB THIPP TRNT
1 1 1 1 1 1 1 1 1 1 1 1 1
UMS WKA
1 1 # for trnL
length(unique(combtab$container[which(combtab$trnL=="trnL")]))[1] 40sort(table(combtab$container[which(combtab$trnL=="trnL")]), decreasing=T) #PNG, YNPBP, UHURU
UHURU BRYCA YNPBP PNG TMBRY BRYBC MADA BRALT HAIDA YHT PANFT DBZN KIFRH
1598 1125 502 496 296 174 164 159 144 89 70 60 51
EDTOL MPIN FOND BBIHP KIFMO SASCH SAFH SAA GTICO KIFAT SABIO IMMEX INDET
50 49 47 33 32 32 24 19 17 11 9 8 8
LIVCA MCT MIPDG LWT CBIHP CBL EMPWG TSA APOSA MKPCH NNOR VASCB FBPL
8 8 7 6 5 4 4 4 3 2 2 2 1
MKTRT
1 # percent of trnL sequences originating from top 5
sum(sort(table(combtab$container[which(combtab$trnL=="trnL")]), decreasing=T)[1:5])/sum(sort(table(combtab$container[which(combtab$trnL=="trnL")]), decreasing=T)) [1] 0.7545079(sort(table(combtab$container[which(combtab$trnL=="trnL")]), decreasing=T)[1:5])/sum(sort(table(combtab$container[which(combtab$trnL=="trnL")]), decreasing=T))
UHURU BRYCA YNPBP PNG TMBRY
0.30015026 0.21130729 0.09429001 0.09316304 0.05559730 # number of specimen records mined from genbank
combtab_mined_from_genbank<- subset (combtab, institution_storing == "Mined from GenBank, NCBI")
head(combtab_mined_from_genbank, 2) X.1 X processid institution_storing phylum_taxID phylum_name
4 11 11 GBITS29777-21 Mined from GenBank, NCBI 12 Magnoliophyta
5 12 12 GBITS29782-21 Mined from GenBank, NCBI 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
4 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
5 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
4 NA <NA> 148601 Aphelandra 801590
5 NA <NA> 148601 Aphelandra 1100819
species_name subspecies_taxID subspecies_name collectiondate_start
4 Aphelandra hylaea NA <NA> NA
5 Aphelandra maximiliana NA <NA> NA
collectiondate_end lat lon coord_source coord_accuracy elev country
4 NA NA NA <NA> NA NA <NA>
5 NA NA NA <NA> NA NA <NA>
province_state region sector exactsite rbcL matK trnL ITS2 multi gb_rbcL
4 <NA> <NA> <NA> <NA> <NA> <NA> <NA> ITS #NAME? <NA>
5 <NA> <NA> <NA> <NA> <NA> <NA> <NA> ITS #NAME? <NA>
gb_matK gb_trnL gb_ITS barcode_count container
4 <NA> <NA> <NA> 1 GBITS
5 <NA> <NA> <NA> 1 GBITS# number of specimens in combtab identified to species
combtab_IDd_to_species <- subset(combtab, species_name != "")
head(combtab_IDd_to_species, 2) X.1 X processid institution_storing phylum_taxID
1 1 1 API120-12 Sri Ramaswamy Memorial University 12
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
1 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
1 Acanthaceae NA <NA> 415894 Peristrophe
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
species_taxID species_name subspecies_taxID subspecies_name
1 494465 Peristrophe bicalyculata NA <NA>
2 632816 Ruellia inflata NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
1 NA NA NA NA <NA> NA
2 NA NA NA NA <NA> NA
elev country province_state region sector exactsite rbcL matK trnL ITS2
1 NA <NA> <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA>
2 NA Brazil Para <NA> <NA> <NA> rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
1 rbcL--- <NA> <NA> <NA> <NA> 1 API
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGI# number of sequences mined from genbank
combtab_mined_from_genbank<- subset (combtab, institution_storing == "Mined from GenBank, NCBI")
head(combtab_mined_from_genbank, 2) X.1 X processid institution_storing phylum_taxID phylum_name
4 11 11 GBITS29777-21 Mined from GenBank, NCBI 12 Magnoliophyta
5 12 12 GBITS29782-21 Mined from GenBank, NCBI 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
4 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
5 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
4 NA <NA> 148601 Aphelandra 801590
5 NA <NA> 148601 Aphelandra 1100819
species_name subspecies_taxID subspecies_name collectiondate_start
4 Aphelandra hylaea NA <NA> NA
5 Aphelandra maximiliana NA <NA> NA
collectiondate_end lat lon coord_source coord_accuracy elev country
4 NA NA NA <NA> NA NA <NA>
5 NA NA NA <NA> NA NA <NA>
province_state region sector exactsite rbcL matK trnL ITS2 multi gb_rbcL
4 <NA> <NA> <NA> <NA> <NA> <NA> <NA> ITS #NAME? <NA>
5 <NA> <NA> <NA> <NA> <NA> <NA> <NA> ITS #NAME? <NA>
gb_matK gb_trnL gb_ITS barcode_count container
4 <NA> <NA> <NA> 1 GBITS
5 <NA> <NA> <NA> 1 GBITScombtab_mined_from_genbank[combtab_mined_from_genbank == ""] <- NA # Convert blank cells to NA
combtab_mined_from_genbank$barcode_count_mined <- rowSums(!is.na(combtab_mined_from_genbank[, c("rbcL", "matK", "trnL", "ITS2")])) # Count the number of barcodes per specimen
total_barcodes_mined <- sum(combtab_mined_from_genbank$barcode_count_mined) # Calculate the total number of barcodes across all rows
#View(combtab_mined_from_genbank)
total_barcodes_mined[1] 199591# number of specimens in combtab with rbcL barcode
combtab_rbcL_barcodes <- subset(combtab, rbcL != "")
head(combtab_rbcL_barcodes, 2) X.1 X processid institution_storing phylum_taxID
1 1 1 API120-12 Sri Ramaswamy Memorial University 12
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
1 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
1 Acanthaceae NA <NA> 415894 Peristrophe
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
species_taxID species_name subspecies_taxID subspecies_name
1 494465 Peristrophe bicalyculata NA <NA>
2 632816 Ruellia inflata NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
1 NA NA NA NA <NA> NA
2 NA NA NA NA <NA> NA
elev country province_state region sector exactsite rbcL matK trnL ITS2
1 NA <NA> <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA>
2 NA Brazil Para <NA> <NA> <NA> rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
1 rbcL--- <NA> <NA> <NA> <NA> 1 API
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGI# number of specimens in combtab with matK barcode
combtab_matK_barcodes <- subset(combtab, matK != "")
head(combtab_matK_barcodes, 2) X.1 X processid institution_storing phylum_taxID
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
3 8 8 EBLF1456-22 Herbarium of South China Botanical Garden 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
3 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
3 Acanthaceae NA <NA> 1142731 Phlogacanthus
species_taxID species_name subspecies_taxID subspecies_name
2 632816 Ruellia inflata NA <NA>
3 1142732 Phlogacanthus colaniae NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
2 NA NA NA NA <NA> NA
3 NA NA NA NA <NA> NA
elev country province_state region sector
2 NA Brazil Para <NA> <NA>
3 220 China Guangxi Chongzuo City <NA>
exactsite rbcL matK trnL ITS2
2 <NA> rbcL matK <NA> ITS
3 Nonggang Protection Station-No.3 Boundary Tablet-L rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGI
3 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 EBLF# number of specimens in combtab with ITS barcode
combtab_ITS2_barcodes <- subset(combtab, ITS2 != "")
head(combtab_ITS2_barcodes, 2) X.1 X processid institution_storing phylum_taxID
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
3 8 8 EBLF1456-22 Herbarium of South China Botanical Garden 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
3 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
3 Acanthaceae NA <NA> 1142731 Phlogacanthus
species_taxID species_name subspecies_taxID subspecies_name
2 632816 Ruellia inflata NA <NA>
3 1142732 Phlogacanthus colaniae NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
2 NA NA NA NA <NA> NA
3 NA NA NA NA <NA> NA
elev country province_state region sector
2 NA Brazil Para <NA> <NA>
3 220 China Guangxi Chongzuo City <NA>
exactsite rbcL matK trnL ITS2
2 <NA> rbcL matK <NA> ITS
3 Nonggang Protection Station-No.3 Boundary Tablet-L rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGI
3 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 EBLF# number of specimens in combtab with trnL barcode
combtab_trnL_barcodes <- subset(combtab, trnL != "")
head(combtab_trnL_barcodes, 2) X.1 X processid
59 78 78 UHURU149-14
60 82 82 UHURU394-14
institution_storing
59 Smithsonian Institution, National Museum of Natural History, United States National Herbarium
60 Smithsonian Institution, National Museum of Natural History, United States National Herbarium
phylum_taxID phylum_name class_taxID class_name order_taxID order_name
59 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
60 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
family_taxID family_name subfamily_taxID subfamily_name genus_taxID
59 148533 Acanthaceae NA <NA> 148750
60 148533 Acanthaceae NA <NA> 306313
genus_name species_taxID species_name subspecies_taxID
59 Justicia 591068 Justicia diclipteroides NA
60 Crossandra 659703 Crossandra massaica NA
subspecies_name collectiondate_start collectiondate_end lat lon
59 <NA> NA NA 0.4810 36.8740
60 <NA> NA NA 0.2956 36.8824
coord_source coord_accuracy elev country province_state region sector
59 <NA> NA 1650 Kenya Rift Valley Laikipia <NA>
60 <NA> NA 1650 Kenya Rift Valley Laikipia <NA>
exactsite rbcL matK trnL ITS2 multi gb_rbcL gb_matK
59 Mpala Research Centre rbcL matK trnL <NA> rbcL-matK-trnL- KM255065 KM255065
60 Mpala Research Centre rbcL matK trnL <NA> rbcL-matK-trnL- JX517979 JX517979
gb_trnL gb_ITS barcode_count container
59 KM255065 <NA> 3 UHURU
60 JX517979 <NA> 3 UHURU# number of genbank accession numbers in combtab
combtab_for_genbank <- combtab
combtab_for_genbank[combtab_for_genbank == ""] <- NA # Convert blank cells to NA
# Count the number of barcodes per specimen (number of non-NA values per row in the specified columns)
combtab_for_genbank$accession_count <- rowSums(!is.na(combtab_for_genbank[, c("gb_rbcL", "gb_matK", "gb_trnL", "gb_ITS")]))
# Calculate the total number of barcodes across all rows (sum of accession_count)
total_accessions <- sum(combtab_for_genbank$accession_count)
# Display the results
#View(combtab_for_genbank)
total_accessions[1] 262920# number of specimens with country listed
combtab_countries <- subset(combtab, country != "")
head(combtab_countries, 2) X.1 X processid institution_storing phylum_taxID
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
3 8 8 EBLF1456-22 Herbarium of South China Botanical Garden 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
3 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
3 Acanthaceae NA <NA> 1142731 Phlogacanthus
species_taxID species_name subspecies_taxID subspecies_name
2 632816 Ruellia inflata NA <NA>
3 1142732 Phlogacanthus colaniae NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
2 NA NA NA NA <NA> NA
3 NA NA NA NA <NA> NA
elev country province_state region sector
2 NA Brazil Para <NA> <NA>
3 220 China Guangxi Chongzuo City <NA>
exactsite rbcL matK trnL ITS2
2 <NA> rbcL matK <NA> ITS
3 Nonggang Protection Station-No.3 Boundary Tablet-L rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGI
3 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 EBLF# number of specimens with lat/long listed
combtab_lat <- subset(combtab, lat != "")
head(combtab_lat, 2) X.1 X processid institution_storing phylum_taxID phylum_name
48 55 55 GBVU4382-13 Mined from GenBank, NCBI 12 Magnoliophyta
49 56 56 GENG1220-15 Gujarat Biodiversity Gene Bank 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
48 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
49 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
48 NA <NA> 554845 Petalidium 554846
49 480072 Thunbergioideae 273749 Thunbergia 317700
species_name subspecies_taxID subspecies_name
48 Petalidium oblongifolium NA <NA>
49 Thunbergia grandiflora NA <NA>
collectiondate_start collectiondate_end lat lon
48 NA NA -28.5536 24.7525
49 NA NA 22.3060 73.2780
coord_source coord_accuracy elev country
48 Coordinates from country centroid NA NA South Africa
49 Garmin etrax 10 NA 130 India
province_state region sector exactsite rbcL matK trnL ITS2 multi
48 <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
49 Gujarat Vodadara <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
48 FJ868916 <NA> <NA> <NA> 1 GBVU
49 KX641599 <NA> <NA> <NA> 1 GENGcombtab_lon <- subset(combtab, lon != "")
head(combtab_lon, 2) X.1 X processid institution_storing phylum_taxID phylum_name
48 55 55 GBVU4382-13 Mined from GenBank, NCBI 12 Magnoliophyta
49 56 56 GENG1220-15 Gujarat Biodiversity Gene Bank 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
48 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
49 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
48 NA <NA> 554845 Petalidium 554846
49 480072 Thunbergioideae 273749 Thunbergia 317700
species_name subspecies_taxID subspecies_name
48 Petalidium oblongifolium NA <NA>
49 Thunbergia grandiflora NA <NA>
collectiondate_start collectiondate_end lat lon
48 NA NA -28.5536 24.7525
49 NA NA 22.3060 73.2780
coord_source coord_accuracy elev country
48 Coordinates from country centroid NA NA South Africa
49 Garmin etrax 10 NA 130 India
province_state region sector exactsite rbcL matK trnL ITS2 multi
48 <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
49 Gujarat Vodadara <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
48 FJ868916 <NA> <NA> <NA> 1 GBVU
49 KX641599 <NA> <NA> <NA> 1 GENG# number of matK sequences
combtab_matK <- subset(combtab, matK == "matK")
head(combtab_matK, 2) X.1 X processid institution_storing phylum_taxID
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
3 8 8 EBLF1456-22 Herbarium of South China Botanical Garden 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
3 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
3 Acanthaceae NA <NA> 1142731 Phlogacanthus
species_taxID species_name subspecies_taxID subspecies_name
2 632816 Ruellia inflata NA <NA>
3 1142732 Phlogacanthus colaniae NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
2 NA NA NA NA <NA> NA
3 NA NA NA NA <NA> NA
elev country province_state region sector
2 NA Brazil Para <NA> <NA>
3 220 China Guangxi Chongzuo City <NA>
exactsite rbcL matK trnL ITS2
2 <NA> rbcL matK <NA> ITS
3 Nonggang Protection Station-No.3 Boundary Tablet-L rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGI
3 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 EBLF# number of matK sequences with lat/long data
combtab_matK_latlong <- subset(combtab_matK, lat != "")
head(combtab_matK_latlong, 2) X.1 X processid
51 58 58 IMDB014-14
52 60 60 KNPA505-09
institution_storing
51 Birla Institute of Technology - Pilani, K.K.Birla Goa Campus
52 University of Johannesburg, Department of Botany and Plant Biotechnology
phylum_taxID phylum_name class_taxID class_name order_taxID order_name
51 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
52 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
family_taxID family_name subfamily_taxID subfamily_name genus_taxID
51 148533 Acanthaceae NA <NA> 390091
52 148533 Acanthaceae NA <NA> 205058
genus_name species_taxID species_name subspecies_taxID
51 Acanthus 641300 Acanthus ilicifolius NA
52 Ruspolia 205059 Ruspolia hypocrateriformis NA
subspecies_name collectiondate_start collectiondate_end lat lon
51 <NA> NA NA 15.5256 73.8753
52 <NA> NA NA -23.9440 31.7250
coord_source coord_accuracy elev country province_state region
51 <NA> NA NA India Goa North Goa
52 <NA> NA 208 South Africa Limpopo <NA>
sector exactsite rbcL
51 Mandovi estuary Chorao Island rbcL
52 Kruger National Park Kruger National Park, Nxanatseni region North, L rbcL
matK trnL ITS2 multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count
51 matK <NA> <NA> rbcL-matK-- DQ028462 DQ028462 <NA> <NA> 2
52 matK <NA> <NA> rbcL-matK-- KY632577 KY632577 <NA> <NA> 2
container
51 IMDB
52 KNPA# number of rbcL sequences
combtab_rbcL <- subset(combtab, rbcL == "rbcL")
head(combtab_rbcL, 2) X.1 X processid institution_storing phylum_taxID
1 1 1 API120-12 Sri Ramaswamy Memorial University 12
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
1 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
1 Acanthaceae NA <NA> 415894 Peristrophe
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
species_taxID species_name subspecies_taxID subspecies_name
1 494465 Peristrophe bicalyculata NA <NA>
2 632816 Ruellia inflata NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
1 NA NA NA NA <NA> NA
2 NA NA NA NA <NA> NA
elev country province_state region sector exactsite rbcL matK trnL ITS2
1 NA <NA> <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA>
2 NA Brazil Para <NA> <NA> <NA> rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
1 rbcL--- <NA> <NA> <NA> <NA> 1 API
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGI# number of rbcL sequences with lat/long data
combtab_rbcL_latlong <- subset(combtab_rbcL, lat != "")
head(combtab_rbcL_latlong, 2) X.1 X processid institution_storing phylum_taxID phylum_name
48 55 55 GBVU4382-13 Mined from GenBank, NCBI 12 Magnoliophyta
49 56 56 GENG1220-15 Gujarat Biodiversity Gene Bank 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
48 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
49 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
48 NA <NA> 554845 Petalidium 554846
49 480072 Thunbergioideae 273749 Thunbergia 317700
species_name subspecies_taxID subspecies_name
48 Petalidium oblongifolium NA <NA>
49 Thunbergia grandiflora NA <NA>
collectiondate_start collectiondate_end lat lon
48 NA NA -28.5536 24.7525
49 NA NA 22.3060 73.2780
coord_source coord_accuracy elev country
48 Coordinates from country centroid NA NA South Africa
49 Garmin etrax 10 NA 130 India
province_state region sector exactsite rbcL matK trnL ITS2 multi
48 <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
49 Gujarat Vodadara <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
48 FJ868916 <NA> <NA> <NA> 1 GBVU
49 KX641599 <NA> <NA> <NA> 1 GENG# number of ITS sequences
combtab_ITS <- subset(combtab, ITS2 == "ITS")
head(combtab_ITS, 2) X.1 X processid institution_storing phylum_taxID
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
3 8 8 EBLF1456-22 Herbarium of South China Botanical Garden 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
3 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
3 Acanthaceae NA <NA> 1142731 Phlogacanthus
species_taxID species_name subspecies_taxID subspecies_name
2 632816 Ruellia inflata NA <NA>
3 1142732 Phlogacanthus colaniae NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
2 NA NA NA NA <NA> NA
3 NA NA NA NA <NA> NA
elev country province_state region sector
2 NA Brazil Para <NA> <NA>
3 220 China Guangxi Chongzuo City <NA>
exactsite rbcL matK trnL ITS2
2 <NA> rbcL matK <NA> ITS
3 Nonggang Protection Station-No.3 Boundary Tablet-L rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGI
3 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 EBLF# number of ITS sequences with lat/long data
combtab_ITS_latlong <- subset(combtab_ITS, lat != "")
head(combtab_ITS_latlong, 2) X.1 X processid institution_storing phylum_taxID
53 62 62 MHPAD3203-10 Area de Conservacion Guanacaste 12
54 65 65 MHPAD537-09 Area de Conservacion Guanacaste 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
53 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
54 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
53 Acanthaceae NA <NA> 148750 Justicia
54 Acanthaceae NA <NA> 148750 Justicia
species_taxID species_name subspecies_taxID subspecies_name
53 317738 Justicia brenesii NA <NA>
54 204815 Justicia arborecens NA <NA>
collectiondate_start collectiondate_end lat lon coord_source
53 NA NA 10.905 -85.279 <NA>
54 NA NA 10.923 -85.465 <NA>
coord_accuracy elev country province_state
53 NA 410 Costa Rica Alajuela
54 NA 1039 Costa Rica Guanacaste
region sector
53 Area de Conservacion Guanacaste Rincon Rainforest
54 Area de Conservacion Guanacaste Sector Cacao
exactsite rbcL matK trnL ITS2
53 Sendero Rincon rbcL matK <NA> ITS
54 Estacion Biologica Cacao, Sendero Arenales rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
53 rbcL-matK--ITS AB114507 AB114507 <NA> AB114507 3 MHPAD
54 rbcL-matK--ITS AF188127 AF188127 <NA> AF188127 3 MHPAD# number of trnL sequences
combtab_trnL <- subset(combtab, trnL == "trnL")
head(combtab_trnL, 2) X.1 X processid
59 78 78 UHURU149-14
60 82 82 UHURU394-14
institution_storing
59 Smithsonian Institution, National Museum of Natural History, United States National Herbarium
60 Smithsonian Institution, National Museum of Natural History, United States National Herbarium
phylum_taxID phylum_name class_taxID class_name order_taxID order_name
59 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
60 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
family_taxID family_name subfamily_taxID subfamily_name genus_taxID
59 148533 Acanthaceae NA <NA> 148750
60 148533 Acanthaceae NA <NA> 306313
genus_name species_taxID species_name subspecies_taxID
59 Justicia 591068 Justicia diclipteroides NA
60 Crossandra 659703 Crossandra massaica NA
subspecies_name collectiondate_start collectiondate_end lat lon
59 <NA> NA NA 0.4810 36.8740
60 <NA> NA NA 0.2956 36.8824
coord_source coord_accuracy elev country province_state region sector
59 <NA> NA 1650 Kenya Rift Valley Laikipia <NA>
60 <NA> NA 1650 Kenya Rift Valley Laikipia <NA>
exactsite rbcL matK trnL ITS2 multi gb_rbcL gb_matK
59 Mpala Research Centre rbcL matK trnL <NA> rbcL-matK-trnL- KM255065 KM255065
60 Mpala Research Centre rbcL matK trnL <NA> rbcL-matK-trnL- JX517979 JX517979
gb_trnL gb_ITS barcode_count container
59 KM255065 <NA> 3 UHURU
60 JX517979 <NA> 3 UHURU# number of ITS sequences with lat/long data
combtab_trnL_latlong <- subset(combtab_trnL, lat != "")
head(combtab_trnL_latlong, 2) X.1 X processid
59 78 78 UHURU149-14
60 82 82 UHURU394-14
institution_storing
59 Smithsonian Institution, National Museum of Natural History, United States National Herbarium
60 Smithsonian Institution, National Museum of Natural History, United States National Herbarium
phylum_taxID phylum_name class_taxID class_name order_taxID order_name
59 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
60 12 Magnoliophyta 41 Magnoliopsida 121216 Lamiales
family_taxID family_name subfamily_taxID subfamily_name genus_taxID
59 148533 Acanthaceae NA <NA> 148750
60 148533 Acanthaceae NA <NA> 306313
genus_name species_taxID species_name subspecies_taxID
59 Justicia 591068 Justicia diclipteroides NA
60 Crossandra 659703 Crossandra massaica NA
subspecies_name collectiondate_start collectiondate_end lat lon
59 <NA> NA NA 0.4810 36.8740
60 <NA> NA NA 0.2956 36.8824
coord_source coord_accuracy elev country province_state region sector
59 <NA> NA 1650 Kenya Rift Valley Laikipia <NA>
60 <NA> NA 1650 Kenya Rift Valley Laikipia <NA>
exactsite rbcL matK trnL ITS2 multi gb_rbcL gb_matK
59 Mpala Research Centre rbcL matK trnL <NA> rbcL-matK-trnL- KM255065 KM255065
60 Mpala Research Centre rbcL matK trnL <NA> rbcL-matK-trnL- JX517979 JX517979
gb_trnL gb_ITS barcode_count container
59 KM255065 <NA> 3 UHURU
60 JX517979 <NA> 3 UHURU# number of overall sequences with lat/long data
combtab_latlong <- subset(combtab, lat != "")
head(combtab_latlong, 2) X.1 X processid institution_storing phylum_taxID phylum_name
48 55 55 GBVU4382-13 Mined from GenBank, NCBI 12 Magnoliophyta
49 56 56 GENG1220-15 Gujarat Biodiversity Gene Bank 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
48 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
49 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
48 NA <NA> 554845 Petalidium 554846
49 480072 Thunbergioideae 273749 Thunbergia 317700
species_name subspecies_taxID subspecies_name
48 Petalidium oblongifolium NA <NA>
49 Thunbergia grandiflora NA <NA>
collectiondate_start collectiondate_end lat lon
48 NA NA -28.5536 24.7525
49 NA NA 22.3060 73.2780
coord_source coord_accuracy elev country
48 Coordinates from country centroid NA NA South Africa
49 Garmin etrax 10 NA 130 India
province_state region sector exactsite rbcL matK trnL ITS2 multi
48 <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
49 Gujarat Vodadara <NA> <NA> rbcL <NA> <NA> <NA> rbcL---
gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
48 FJ868916 <NA> <NA> <NA> 1 GBVU
49 KX641599 <NA> <NA> <NA> 1 GENGtable(combtab$trnL)
trnL
5324 levels(factor(combtab$trnL))[1] "trnL"Figure 1B-C. Plots for number of specimens across lat and lon for each barcode
# round lat and lon to nearest one degree
latr <-round(combtab$lat)
#latr
lonr <- round(combtab$lon)
#lonr
# create rounded data frame
combtab_rounded <- combtab %>% dplyr::mutate(dplyr::across(c('lat', 'lon'), round, 0))
head(combtab_rounded, 2) X.1 X processid institution_storing phylum_taxID
1 1 1 API120-12 Sri Ramaswamy Memorial University 12
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
1 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
1 Acanthaceae NA <NA> 415894 Peristrophe
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
species_taxID species_name subspecies_taxID subspecies_name
1 494465 Peristrophe bicalyculata NA <NA>
2 632816 Ruellia inflata NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
1 NA NA NA NA <NA> NA
2 NA NA NA NA <NA> NA
elev country province_state region sector exactsite rbcL matK trnL ITS2
1 NA <NA> <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA>
2 NA Brazil Para <NA> <NA> <NA> rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
1 rbcL--- <NA> <NA> <NA> <NA> 1 API
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGIcombtab_rounded_38 <- subset(combtab_rounded, lat == 38)
head(combtab_rounded_38, 2) X.1 X processid institution_storing phylum_taxID phylum_name
366 507 507 GBVY3489-14 Mined from GenBank, NCBI 12 Magnoliophyta
1312 2051 2051 GBVU4183-13 Mined from GenBank, NCBI 12 Magnoliophyta
class_taxID class_name order_taxID order_name family_taxID family_name
366 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
1312 41 Magnoliopsida 121216 Lamiales 148533 Acanthaceae
subfamily_taxID subfamily_name genus_taxID genus_name species_taxID
366 480072 Thunbergioideae 273749 Thunbergia 601895
1312 264310 Acanthoideae 415904 Andrographis 415905
species_name subspecies_taxID subspecies_name
366 Thunbergia coccinea NA <NA>
1312 Andrographis paniculata NA <NA>
collectiondate_start collectiondate_end lat lon
366 NA NA 38 105
1312 NA NA 38 105
coord_source coord_accuracy elev country
366 Coordinates from country centroid NA NA China
1312 Coordinates from country centroid NA NA China
province_state region sector exactsite rbcL matK trnL ITS2
366 <NA> Yunnan, Mengsong <NA> <NA> rbcL <NA> <NA> <NA>
1312 <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA>
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
366 rbcL--- KR737446 <NA> <NA> <NA> 1 GBVY
1312 rbcL--- <NA> <NA> <NA> <NA> 1 GBVU# Plot longitude
marker_count_by_long_plot2 <- ggplot(combtab_rounded, aes(x = lon)) +
theme_classic() +
geom_bar(stat = "count", position = "dodge", fill = "gray") +
scale_x_continuous(breaks = seq(-180, 180, 60), limits = c(-180, 180)) + scale_y_continuous(breaks = seq(0, 5000, 1000), limits = c(0, 5000)) +
geom_line(data = subset(combtab_rounded, matK == "matK"), aes(x = lon), stat = "count", col = "#62BAAC", lwd = 0.4) +
geom_line(data = subset(combtab_rounded, rbcL == "rbcL"), aes(x = lon), stat = "count", col = "#EDA247", lwd = 0.4) +
geom_line(data = subset(combtab_rounded, trnL == "trnL"), aes(x = lon), stat = "count", col = "#006B5F", lwd = 0.4) +
geom_line(data = subset(combtab_rounded, ITS2 == "ITS"), aes(x = lon), stat = "count", col = "#703900", lwd = 0.4) +
ylab("") + xlab("") #+ theme(panel.background = element_blank(), axis.text = element_blank())
marker_count_by_long_plot2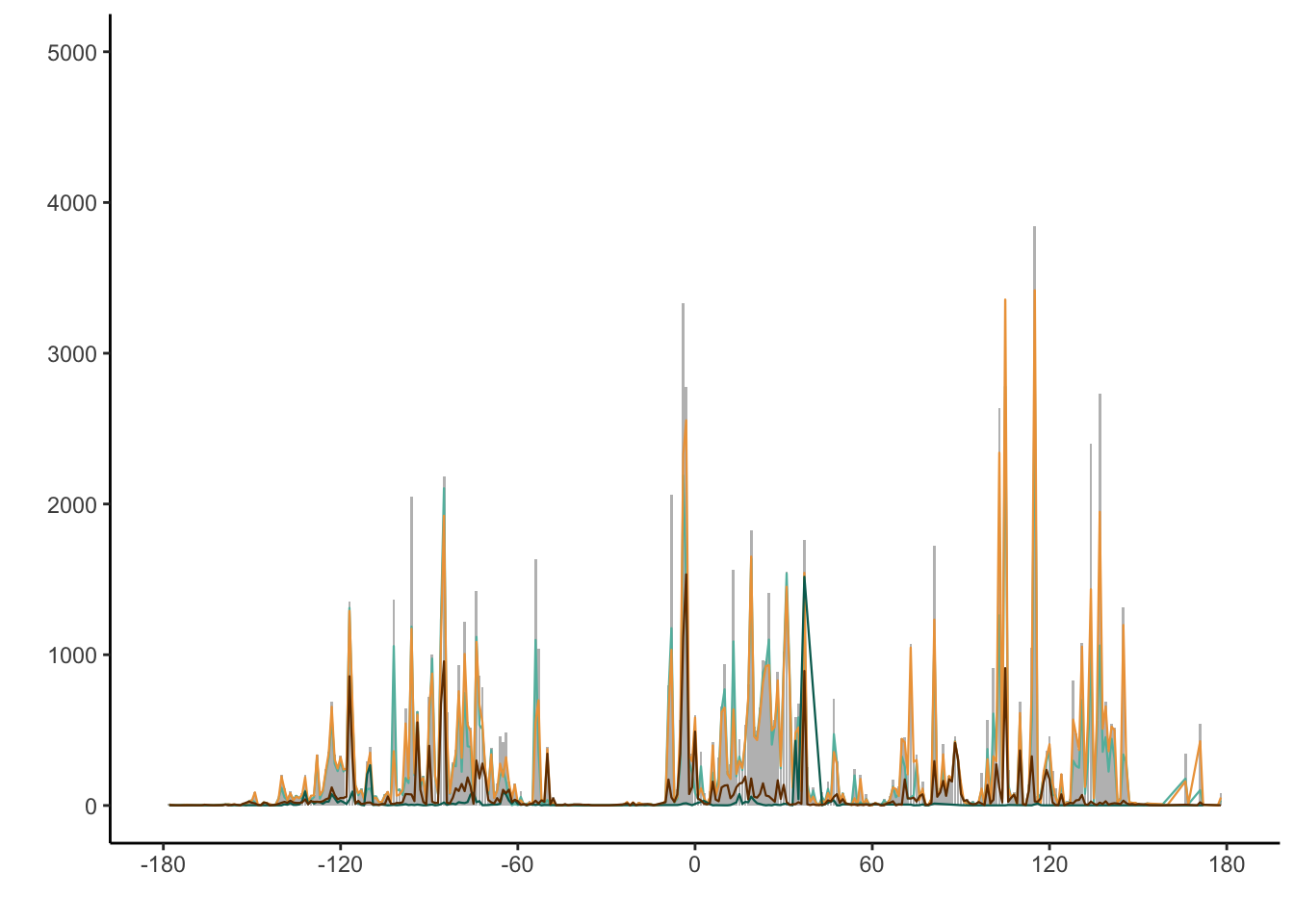
#ggsave("marker_count_by_long_plot2_20240624.pdf", marker_count_by_long_plot2, width = 13, height = 5, unit = "cm")
# Plot latitude
marker_count_by_lat_plot_alllines <- ggplot() +
theme_classic() +
scale_x_continuous(breaks = seq(-90, 90, 45), limits = c(-90, 90)) + scale_y_continuous(breaks = seq(0, 5500, 2500), limits = c(0, 5500)) +
geom_area(data = combtab_rounded, aes(x = lat), stat = "count", fill = "lightgray", lwd = 0.4) +
geom_line(data = subset(combtab_rounded, matK == "matK"), aes(x = lat), stat = "count", col = "#62BAAC", lwd = 0.4) +
geom_line(data = subset(combtab_rounded, rbcL == "rbcL"), aes(x = lat), stat = "count", col = "#EDA247", lwd = 0.4) +
geom_line(data = subset(combtab_rounded, trnL == "trnL"), aes(x = lat), stat = "count", col = "#006B5F", lwd = 0.4)+
geom_line(data = subset(combtab_rounded, ITS2 == "ITS"), aes(x = lat), stat = "count", col = "#703900", lwd = 0.4)+
ylab("") + xlab("") + theme(panel.background = element_blank(), axis.text = element_blank())
marker_count_by_lat_plot_alllines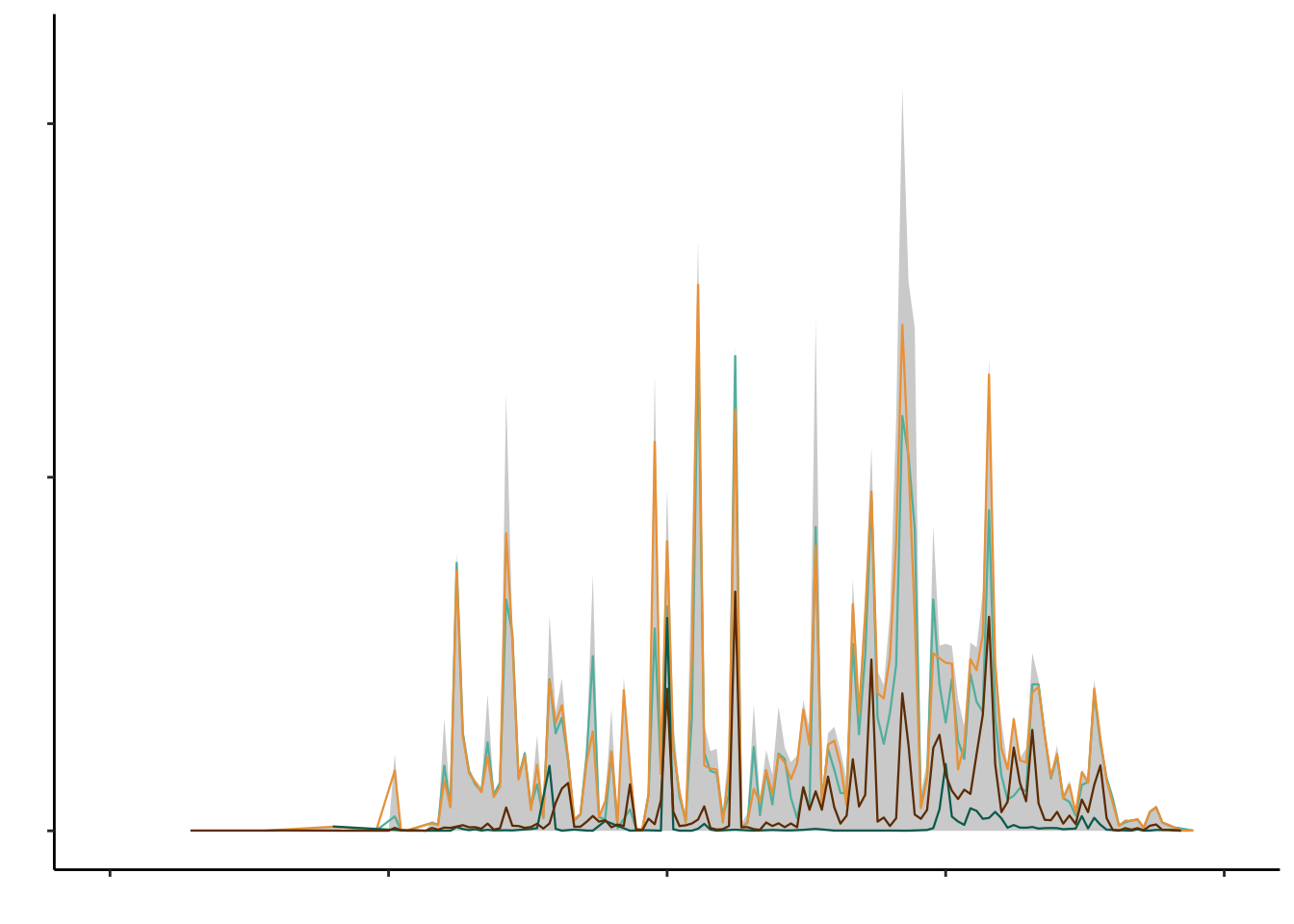
#ggsave("marker_count_by_lat_plot_alllines_20240626_2.pdf", marker_count_by_lat_plot_alllines, width = 8.6, height = 5, unit = "cm")combtab_matK <- sum(combtab$matK == "matK")
combtab_rbcL <- combtab %>% subset(rbcL == "rbcL")
head(combtab_rbcL, 2) X.1 X processid institution_storing phylum_taxID
1 1 1 API120-12 Sri Ramaswamy Memorial University 12
2 2 2 CANGI002-17 Museu Paraense Emilio Goeldi 12
phylum_name class_taxID class_name order_taxID order_name family_taxID
1 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
2 Magnoliophyta 41 Magnoliopsida 121216 Lamiales 148533
family_name subfamily_taxID subfamily_name genus_taxID genus_name
1 Acanthaceae NA <NA> 415894 Peristrophe
2 Acanthaceae 264310 Acanthoideae 148534 Ruellia
species_taxID species_name subspecies_taxID subspecies_name
1 494465 Peristrophe bicalyculata NA <NA>
2 632816 Ruellia inflata NA <NA>
collectiondate_start collectiondate_end lat lon coord_source coord_accuracy
1 NA NA NA NA <NA> NA
2 NA NA NA NA <NA> NA
elev country province_state region sector exactsite rbcL matK trnL ITS2
1 NA <NA> <NA> <NA> <NA> <NA> rbcL <NA> <NA> <NA>
2 NA Brazil Para <NA> <NA> <NA> rbcL matK <NA> ITS
multi gb_rbcL gb_matK gb_trnL gb_ITS barcode_count container
1 rbcL--- <NA> <NA> <NA> <NA> 1 API
2 rbcL-matK--ITS <NA> <NA> <NA> <NA> 3 CANGImatK <- subset(combtab_rounded, matK == "matK")
rbcL <- subset(combtab_rounded, rbcL == "rbcL")
trnL <- subset(combtab_rounded, trnL == "trnL")
ITS <- subset(combtab_rounded, ITS2 == "ITS") Figure 1D. Plot MAT and MAP points with 95% confidence intervals
To run this next section, you’ll need to download temperature and precipitation data from [Worldclim](“https://www.worldclim.org/data/worldclim21.html). Download both the tavg 10m and prec 10m datasets, unzip, and add to your ./data folder at the root of the MolEco-MEC-24-1288 repository (Fick and Hijmans 2017).
#####
# Worldclim connect lat lon specimen records (rounded to nearest degree and dereplicated) with temperature and precip
#####
# dereplicate sites rounded to nearest degree
sitepix<-cbind(lonr, latr)
sitepix<-unique(sitepix)
# load worldclim data
# use 10 min for testing download directly from worldclim
t.mean.files <- list.files("../data/wc2.1_10m_tavg/", ".tif", full.names=TRUE)
p.mean.files <- list.files("../data/wc2.1_10m_prec/", ".tif", full.names=TRUE)
# temperature (using the raster package)
t.mean <- stack(t.mean.files)
t.mean <-mean(t.mean)
# make a 'samples' data frame to extract temp values from barcode localities
samples <- data.frame(processid=seq(1,nrow(sitepix)), lon=sitepix[,1], lat=sitepix[,2], row.names="processid")
head(samples, 2) lon lat
1 NA NA
2 25 -29samples <- samples[which(samples$lat!="NA"),]
head(samples, 2) lon lat
2 25 -29
3 73 22# Extract data from RasterLayer
temp.data <- raster::extract(t.mean, samples)
head(temp.data, 2)[1] 17.8276 27.2780sampletempMeans <- temp.data
length(which(is.na(sampletempMeans))) [1] 393# precipitation
p.mean <- stack(p.mean.files)
p.mean <-sum(p.mean)/10 # divide by 10 to convert from mm to cm
# Extract data from RasterLayer
precip.data <- raster::extract(p.mean, samples)
head(precip.data)[1] 43.6 88.5 53.2 414.5 54.9 275.5sampleprecipMeans <- precip.data
length(which(is.na(sampleprecipMeans))) [1] 393# plot Figure 1d
# get random points to plot as background
set.seed(20140816)
nrand<-100000
randomtemp <- sampleRandom(mean(t.mean), nrand, xy=T)
randomprecip <- extract(mean(p.mean), randomtemp[,1:2])
randomtempMeans <- randomtemp[,3]
randomprecipMeans <- randomprecip
randompoints <- cbind(randomtempMeans, randomprecipMeans)
# find the upper / lower 95 percentile for temp and precip to plot based on BOLD samples
temp95 <- quantile(sampletempMeans, c(0.05), na.rm=T)
precip95 <- quantile(sampleprecipMeans, c(0.95), na.rm=T)
# Plot figure 1D
plot(randomprecipMeans ~ randomtempMeans, xlim=rev(range(randomtempMeans, na.rm=T)), pch=16, col="gray", ylab="Mean annual precipitation (cm)", xlab="mean annual temperature (°C)", type="n", ylim=c(0, max(c(max(randomprecipMeans), max(sampleprecipMeans, na.rm=T)))))
pointcols<-rep("gray", nrand)
pointcols[which(randompoints[,1]<=temp95)]<-"darkblue"
pointcols[which(randompoints[,2]>=precip95)]<-"darkgreen"
pointcols[which(randompoints[, 1] <= temp95 & randompoints[, 2] >= precip95)] <- "orange"
points(randompoints[,1], randompoints[,2], pch=16, col= pointcols)
points(x=sampletempMeans, y=sampleprecipMeans, pch=3, col=alpha("black", 0.2))
abline(h=precip95, v=temp95, lty=2, col="black", lwd=2)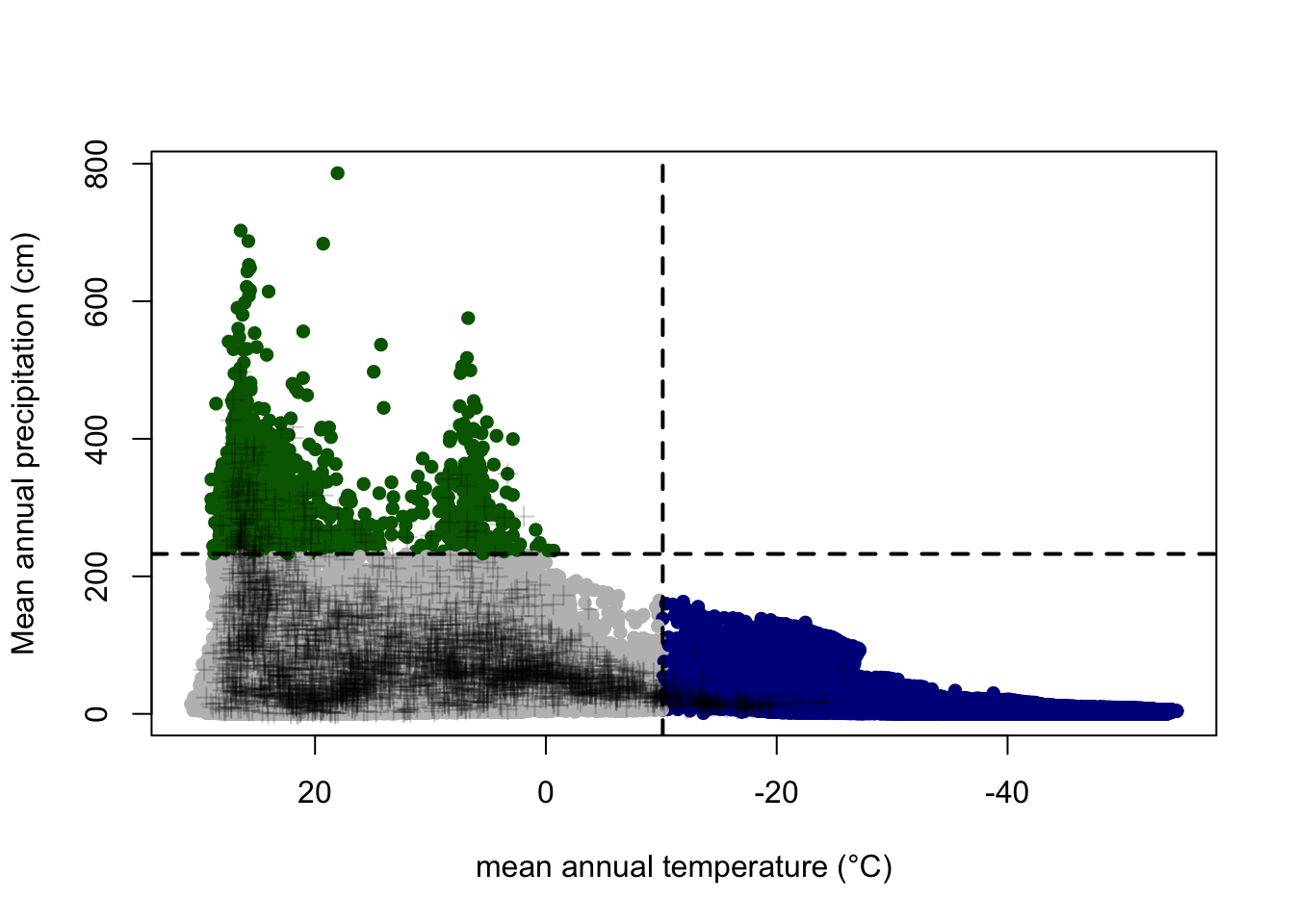
Figure 1A. Map: Geography of plant barcodes.
# map for places inside or outside climate envelopes
# make raster by thresholds
threshtemp <- t.mean
values(threshtemp) <- as.numeric(values(threshtemp<as.numeric(temp95)))
quantile(values(p.mean), na.rm=T) 0% 25% 50% 75% 100%
0.0 11.0 33.8 68.7 1119.1 threshprecip <- p.mean
values(threshprecip) <- as.numeric(values(threshprecip>as.numeric(precip95)))
class(threshprecip)[1] "RasterLayer"
attr(,"package")
[1] "raster"# create ggplot-usable world map
world <- sf::st_as_sf(map("world", plot = FALSE, fill = TRUE))
# transform all features to longlat and WGS84 - first, world
st_crs(world)Coordinate Reference System:
User input: +proj=longlat +ellps=clrk66 +no_defs +type=crs
wkt:
GEOGCRS["unknown",
DATUM["Unknown based on Clarke 1866 ellipsoid",
ELLIPSOID["Clarke 1866",6378206.4,294.978698213898,
LENGTHUNIT["metre",1,
ID["EPSG",9001]]]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8901]],
CS[ellipsoidal,2],
AXIS["longitude",east,
ORDER[1],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]],
AXIS["latitude",north,
ORDER[2],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]]]world <- st_transform(world,CRS("+proj=longlat +datum=WGS84")) # the goal here is to choose a projection and datum that every spatial object we add to the map will be converted to before plotting
st_crs(world) # check coordinate system to ensure projection workedCoordinate Reference System:
User input: GEOGCRS["unknown",
DATUM["World Geodetic System 1984",
ELLIPSOID["WGS 84",6378137,298.257223563,
LENGTHUNIT["metre",1]],
ID["EPSG",6326]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8901]],
CS[ellipsoidal,2],
AXIS["longitude",east,
ORDER[1],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]],
AXIS["latitude",north,
ORDER[2],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]]]
wkt:
GEOGCRS["unknown",
DATUM["World Geodetic System 1984",
ELLIPSOID["WGS 84",6378137,298.257223563,
LENGTHUNIT["metre",1]],
ID["EPSG",6326]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8901]],
CS[ellipsoidal,2],
AXIS["longitude",east,
ORDER[1],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]],
AXIS["latitude",north,
ORDER[2],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]]]world <- fortify(world) # re-examine features
class(world)[1] "sf" "data.frame"# transform all features to longlat and WGS84 - next, threshtemp
st_crs(threshtemp)Coordinate Reference System:
User input: +proj=longlat +datum=WGS84 +no_defs
wkt:
GEOGCRS["unknown",
DATUM["World Geodetic System 1984",
ELLIPSOID["WGS 84",6378137,298.257223563,
LENGTHUNIT["metre",1]],
ID["EPSG",6326]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8901]],
CS[ellipsoidal,2],
AXIS["longitude",east,
ORDER[1],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]],
AXIS["latitude",north,
ORDER[2],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]]]newthreshtemp <- " +proj=longlat +datum=WGS84"
# simplest approach
threshtemp <- projectRaster(threshtemp, crs=newthreshtemp)
st_crs(threshtemp) # check coordinate system to ensure projection workedCoordinate Reference System:
User input: +proj=longlat +datum=WGS84 +no_defs
wkt:
GEOGCRS["unknown",
DATUM["World Geodetic System 1984",
ELLIPSOID["WGS 84",6378137,298.257223563,
LENGTHUNIT["metre",1]],
ID["EPSG",6326]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8901]],
CS[ellipsoidal,2],
AXIS["longitude",east,
ORDER[1],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]],
AXIS["latitude",north,
ORDER[2],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]]]threshtemp #re-examine featuresclass : RasterLayer
dimensions : 1078, 2156, 2324168 (nrow, ncol, ncell)
resolution : 0.167, 0.167 (x, y)
extent : -180, 180.052, -90.026, 90 (xmin, xmax, ymin, ymax)
crs : +proj=longlat +datum=WGS84 +no_defs
source : memory
names : layer
values : 0, 1 (min, max)# transform all features to longlat and WGS84 - next, threshprecip
st_crs(threshprecip)Coordinate Reference System:
User input: +proj=longlat +datum=WGS84 +no_defs
wkt:
GEOGCRS["unknown",
DATUM["World Geodetic System 1984",
ELLIPSOID["WGS 84",6378137,298.257223563,
LENGTHUNIT["metre",1]],
ID["EPSG",6326]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8901]],
CS[ellipsoidal,2],
AXIS["longitude",east,
ORDER[1],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]],
AXIS["latitude",north,
ORDER[2],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]]]newthreshprecip <- " +proj=longlat +datum=WGS84"
# simplest approach
threshprecip <- projectRaster(threshprecip, crs=newthreshprecip)
st_crs(threshprecip) # check coordinate system to ensure projection workedCoordinate Reference System:
User input: +proj=longlat +datum=WGS84 +no_defs
wkt:
GEOGCRS["unknown",
DATUM["World Geodetic System 1984",
ELLIPSOID["WGS 84",6378137,298.257223563,
LENGTHUNIT["metre",1]],
ID["EPSG",6326]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8901]],
CS[ellipsoidal,2],
AXIS["longitude",east,
ORDER[1],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]],
AXIS["latitude",north,
ORDER[2],
ANGLEUNIT["degree",0.0174532925199433,
ID["EPSG",9122]]]]threshprecip # re-examine featuresclass : RasterLayer
dimensions : 1078, 2156, 2324168 (nrow, ncol, ncell)
resolution : 0.167, 0.167 (x, y)
extent : -180, 180.052, -90.026, 90 (xmin, xmax, ymin, ymax)
crs : +proj=longlat +datum=WGS84 +no_defs
source : memory
names : layer
values : 0, 1 (min, max)# Create a data frame for ggplot purposes - threshtemp
threshtemp_df <- as.data.frame(threshtemp, xy = TRUE, na.rm = TRUE)
head(threshtemp_df, 2) x y layer
80618 -38.8015 83.7375 0
80619 -38.6345 83.7375 0threshtemp_df_subset <- subset(threshtemp_df, layer > 0)
# Create a data frame for ggplot purposes - threshprecip
threshprecip_df <- as.data.frame(threshprecip, xy = TRUE, na.rm = TRUE)
head(threshprecip_df, 2) x y layer
80618 -38.8015 83.7375 0
80619 -38.6345 83.7375 0threshprecip_df_subset <- subset(threshprecip_df, layer > 0 )# Plot geography of plant DNA barcodes
#ggplot wrapper of map()
world_map <- map_data("world")
ggmap4 <- ggplot() + theme_classic() +
geom_map(dat = world_map, map = world_map, aes(map_id = region), fill = "white", color = "#7f7f7f", size = 0.25) +
geom_raster(data = threshtemp_df, aes(x = x, y = y, fill = layer), alpha = 0.9) +
scale_fill_gradient(low = "transparent", high = "darkblue", aesthetics = 'fill') +
annotate(geom="raster", x=threshprecip_df_subset$x, y=threshprecip_df_subset$y, alpha=.9,
fill = scales::colour_ramp(c("transparent","darkgreen"))(threshprecip_df_subset$layer)) + geom_point(data = combtab, aes(x = lon, y = lat), pch = 3, size = 0.05, stroke = 0.25, color = alpha("darkgray", 0.9)) + ylab("") + xlab("") + theme(legend.position = "none") + scale_y_continuous(breaks=seq(-90,90,45), limits = c(-90, 90)) + scale_x_continuous(breaks=seq(-180,180,60), limits = c(-180, 180)) + ylab("") + xlab("")+ theme(panel.background = element_blank(), axis.text = element_blank()) +
geom_hline(yintercept = 0, color = "black", size = 0.5) + # Equator line
geom_hline(yintercept = 23.5, linetype = "dashed", color = "black", size = 0.5) + # Tropic of Cancer
geom_hline(yintercept = -23.5, linetype = "dashed", color = "black", size = 0.5) # Tropic of Capricorn
ggmap4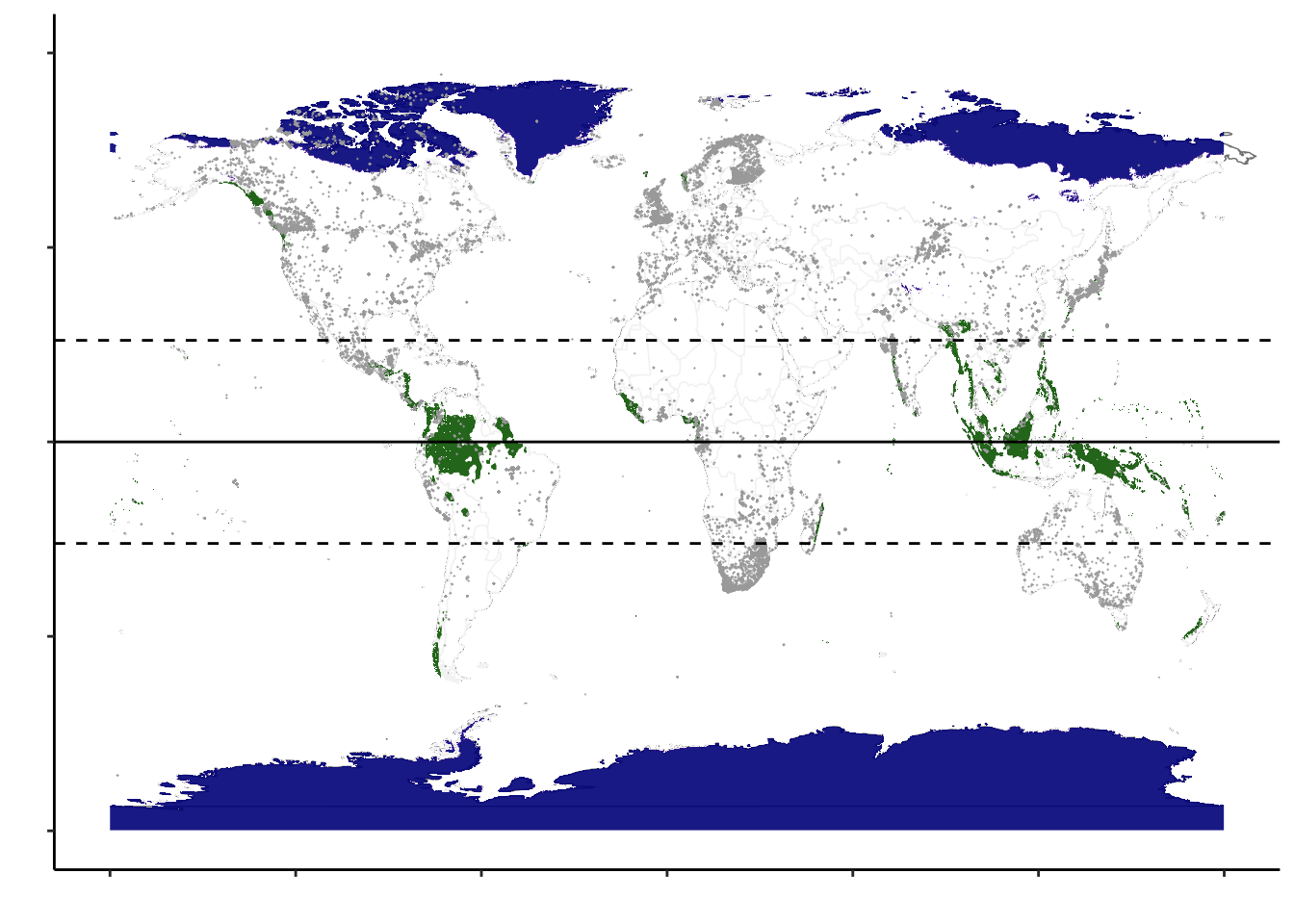
#ggsave("ggmap4_20240626_1.pdf", ggmap4, width = 13, height = 8.6, unit = "cm")